|
3/26/2023 0 Comments Irapp gscaasEight distinct indel variations were detected in the chloroplast genome alignment of 24 Dioscorea species. 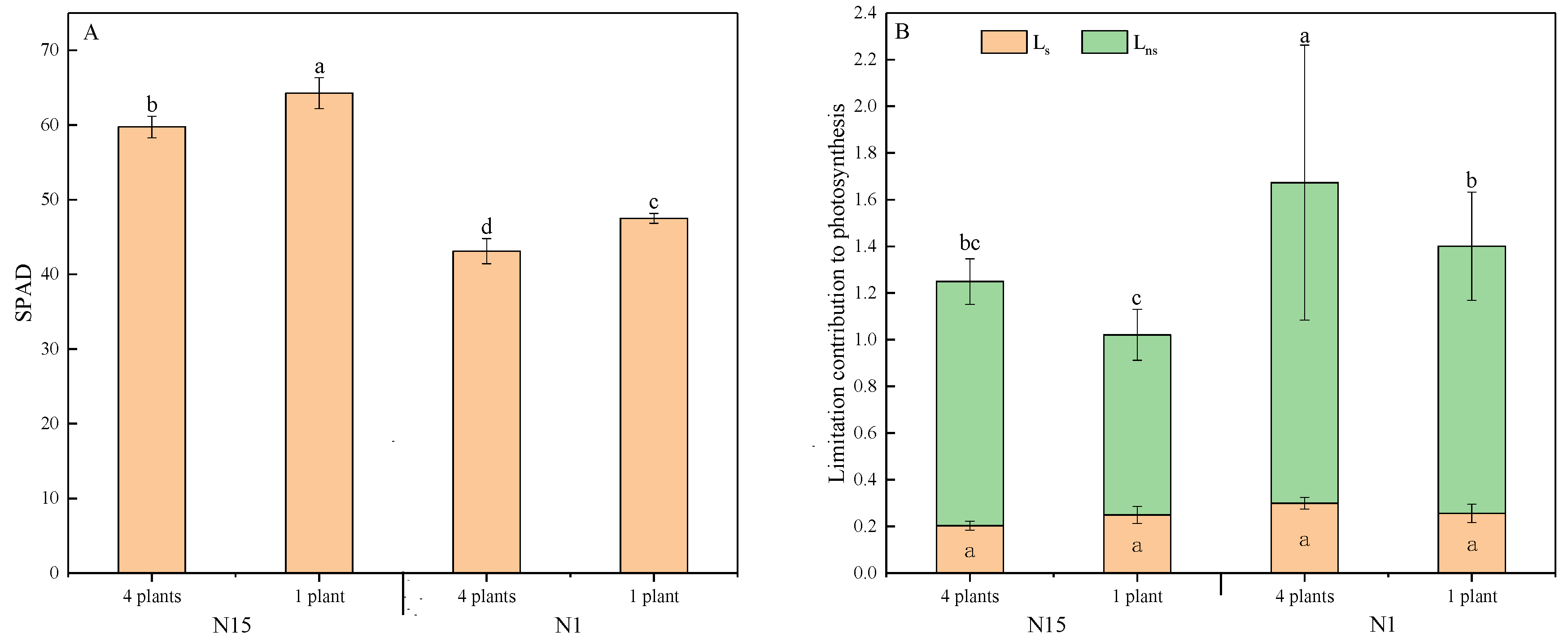
Genes adjacent to the junction borders were similar in all species analyzed. Only mono-, di-, and trinucleotides were present in the genome. The overall GC content for all four species was 37%. A total of 113 genes were predicted, including 79 protein-coding sequences (CDS), 30 tRNA, and four rRNA genes. The chloroplast genomes of Dioscorea brevipetiolata, D. This study contributes to further understanding the evolutionary process and the biodiversity conservation of C. nipponicum is likely introduced through the north Eurasian root, not the mainland, and evolved independently in Ulleung Island. We found 833 polymorphic sites and eight highly variable regions in chloroplast genomes of six Cirsium species by calculating nucleotide diversity, as well as 18 specific variable regions distinguished C. The chloroplast genome was 152,586 bp, encoding 133 genes consisting of 8 rRNA genes, 37 tRNA genes, and 88 protein-coding genes. nipponicum and reconstructed the phylogenetic relationships within the genus Cirsium. 
We thus assembled the complete chloroplast of C. nipponicum, there is not much genomic information to estimate it. Although many researchers have questioned the origin and evolution of C. Unlike other Cirsium in Korea, Cirsium nipponicum (Island thistle) is distributed only on Ulleung Island, a volcanic island off the east coast of the Korean Peninsula, and a unique thistle with none or very small thorns. This study may provide useful information for genetic diversity analysis and molecular identification for Bidens species. Moreover, mutation hotspot analysis and in silico PCR analysis indicated that inter-genic regions of ndhD-ccsA, ndhI-ndhG, ndhF-rpl32, trnL_UAG-rpl32, ndhE-psaC, matK-rps16, rps2-atpI, cemA-petA, petN-psbM were candidate markers of molecular identification for Bidens plants. Phylogenetic analysis revealed that 3 Bidens plants clustered together and further formed molophyletic clade with other Bidens species, indicating Bidens plants might be under radiation adaptive selection to the changing environment world-widely. It seems that ndhE, ndhF, ndhG, and rpl32 from the Bidens plants were under positive selection while the majority of chloroplast genes were under purifying selection. Gene loss of clpP and psbD, IR expansion and contraction were detected in these Bidens plants. Similar number of SSRs and long repeats were found in Bidens, wherein mononucleotide repeats (A/T), forward and palindromic repeats were the most in abundance. The chloroplast genomes are in typical quadripartite structure with two inverted repeat regions separating a large single copy region and a small single copy region, and ranged from 151,599 to 154,478 bp in length. radiata) have been sequenced, assembled and annotated in this study to distinguish their molecular characterization and phylogenetic relationships. Source codes and the online suit available or genomes for 3 Bidens plants endemic to China (Bidens bipinnata Linn., Bidens pilosa Linn., and Bidens alba var. The software was tested using over a hundred embryophyte chloroplast genomes and in all cases a reliable output was obtained. The input of the program is an annotation GenBank (.gb) file, the accession or GI number of the sequence or a DOGMA output file. We introduce a website for easier use of the program as well as R source code of the software to be used in case of preferences to be changed and integrated into personal pipelines. The software and its dependent libraries are fully coded in R and the reflected plot is scaled up to realistic size of nucleotide base pairs in the vicinity of the junction sites. It allows the users to depict the genetic architecture of up to ten chloroplast genomes in the vicinity of the sites connecting the inverted repeats to the short and long single copy regions. Here we announce a new visualization tool that is specifically designed for chloroplast genomes. These tools can provide a deeper insight into the genomic structure of the organism. 
Genome plotting is performed using a wide range of visualizations tools each with emphasis on a different informative dimension of the genome.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
